Sull’importanza di costruire e valutare le strategie di screening tenendo conto del contesto epidemico
Publication date: 14/12/2020 – E&P Code: repo.epiprev.it/2068
Authors: Michela Baccini1
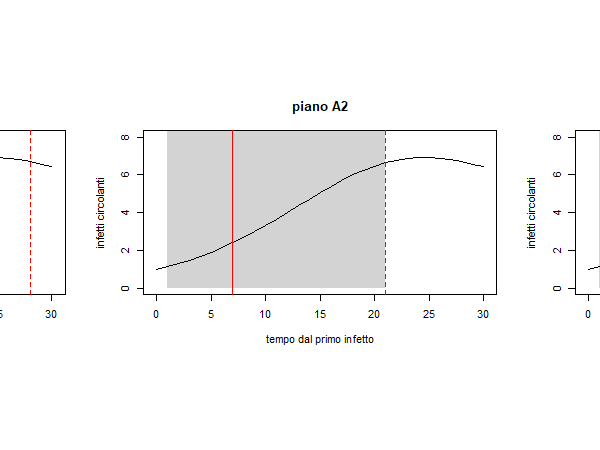
Abstract: Nel predisporre piani di screening per la prevenzione e il contenimento del contagio da SARS‐coV‐2 si dovrebbe tenere conto del contesto epidemico e in particolare del fatto che i soggetti infetti non individuati sono potenzialmente capaci di trasmettere la malattia e che l’infezione si diffonde per focolai, creando dei cluster nella popolazione. A titolo esemplificativo, in questa nota si confrontano due possibili strategie di screening in ambito scolastico, ipotizzando diversi scenari di evoluzione del contagio all’interno di una classe di 20 alunni a partire da un infetto iniziale. Le strategie messe a confronto sono di sue tipi: una procedura basata su test a tappeto ripetuti ogni 15 giorni e una procedura che, suddivisa la classe in due sottogruppi, li sottopone a test a settimane alterne. Quest’ultima strategia sembrerebbe garantire risultati migliori soprattutto quando la trasmissione del contagio entro la classe è più elevata. L’approccio di valutazione proposto può essere adottato anche per confrontare tra loro piani di screening più complessi di quelli qui presi in considerazione.
More >